Host-Pathogen Interactions
Viral Integration ♦ Viroid RNA
Retroviral Integration
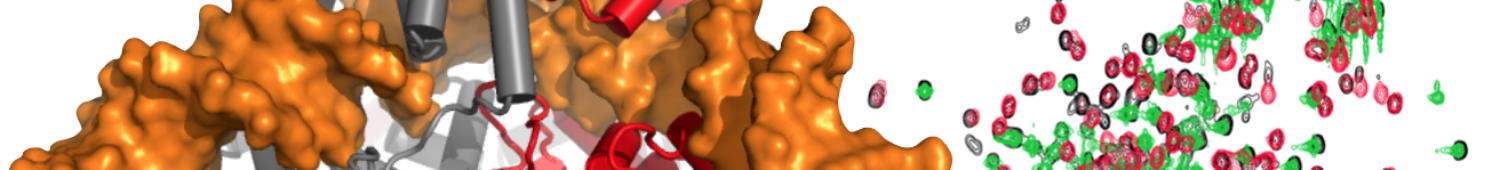
 Retroviral integration facilitated by interactions between retroviral integrases and a series of host-encoded proteins. In collaboration with the laboratory of Mamuka Kvaratskhelia we are using NMR spectroscopy and other structural tools to understand the structural basis for these interactions.
Retroviral integration facilitated by interactions between retroviral integrases and a series of host-encoded proteins. In collaboration with the laboratory of Mamuka Kvaratskhelia we are using NMR spectroscopy and other structural tools to understand the structural basis for these interactions.
Viroid RNA
The Potato Spindle Tuber Viroid (PSTVd) is a closed, circular RNA of only 359 nucleotides in length that features conserved double-stranded and loop regions. PSTVd does not encode any proteins and yet is able to replicate and spread systematically within an infected host plant thanks to conserved structural motifs that are thought to interact with necessary cellular factors. The Biao Ding lab has used genome-wide mutagenesis regions of PSTVd critical for replication (R) or cell-to-cell trafficking (T). We are using NMR spectroscopy to determine the structures of some of these conserved structural features in order to provide a structural foundation for understanding how they interact with host factors in order to carry out their biological functions.