“One of the great things about books is sometimes there are some fantastic pictures.” George W. Bush, Jan. 3, 2000
A
A
The Ohio State University
2024
59. DB Heisler, E Kudryashova, R Hitt, B Williams, M Dziejman, JS Gunn, DS Kudryashov bioRxiv, (2024).07. 01.601609
Antagonistic effects of actin-specific toxins on Salmonella Typhimurium invasion into mammalian cells
Competition between bacterial species is a major factor shaping microbial communities. In this work, we explored the hypothesis that competition between bacterial pathogens can be mediated through antagonistic effects of bacterial effector proteins on host systems, particularly the actin cytoskeleton. Using Salmonella Typhimurium invasion into cells as a model, we demonstrate that invasion is inhibited if the host actin cytoskeleton is disturbed by any of the four tested actin-specific toxins: Vibrio cholerae MARTX actin crosslinking and Rho GTPase inactivation domains (ACD and RID, respectively), TccC3 from Photorhabdus luminescens, and Salmonella`s own SpvB. We noticed that ACD, being an effective inhibitor of tandem G-actin binding assembly factors, is likely to inhibit the activity of another Vibrio effector, VopF. In reconstituted actin polymerization assays confirmed by live-cell microscopy, we confirmed that ACD potently halted the actin nucleation and pointed-end elongation activities of VopF, revealing competition between these two V. cholerae effectors. Together, the results suggest bacterial effectors from different species that target the same host machinery or proteins may represent an effective but largely overlooked mechanism of indirect bacterial competition in host-associated microbial communities. Whether the proposed inhibition mechanism involves the actin cytoskeleton or other host cell compartments, such inhibition deserves investigation and may contribute to a documented scarcity of human enteric co-infections by different pathogenic bacteria.
58. Niedzialkowska E, Runyan LA, Kudryashova E, Egelman EH, Kudryashov DS. Structure. 2024 Jun 6;32(6):725-738.e8. doi: 10.1016/j.str.2024.02.022. Epub 2024 Mar 21. PMID: 38518780
Stabilization of F-actin by Salmonella effector SipA resembles the structural effects of inorganic phosphate and phalloidin.
Entry of Salmonella into host enterocytes relies on its pathogenicity island 1 effector SipA. We found that SipA binds to F-actin in a 1:2 stoichiometry with sub-nanomolar affinity. A cryo-EM reconstruction revealed that SipA’s globular core binds at the groove between actin strands, whereas the extended C-terminal arm penetrates deeply into the inter-strand space, stabilizing F-actin from within. The unusually strong binding of SipA is achieved by a combination of fast association via the core and very slow dissociation dictated by the arm. Similar to Pi, BeF3, and phalloidin, SipA potently inhibited actin depolymerization by actin depolymerizing factor (ADF)/cofilin, which correlated with increased filament stiffness, supporting the hypothesis that F-actin’s mechanical properties contribute to the recognition of its nucleotide state by protein partners. The remarkably strong binding to F-actin maximizes the toxin’s effects at the injection site while minimizing global influence on the cytoskeleton and preventing pathogen detection by the host cell.
2023
57. Zhang Q, Wan M, Kudryashova E, Kudryashov DS, Mao Y. PLoS Pathog. (2023) Jul 18;19(7):e1011512. doi: 10.1371/journal.ppat.1011512. eCollection 2023 Jul. PMID: 37463171
Membrane-dependent actin polymerization mediated by the Legionella pneumophila effector protein MavH.
L. pneumophila propagates in eukaryotic cells within a specialized niche, the Legionella-containing vacuole (LCV). The infection process is controlled by over 330 effector proteins delivered through the type IV secretion system. In this study, we report that the Legionella MavH effector localizes to endosomes and remodels host actin cytoskeleton in a phosphatidylinositol 3-phosphate (PI(3)P) dependent manner when ectopically expressed. We show that MavH recruits host actin capping protein (CP) and actin to the endosome via its CP-interacting (CPI) motif and WH2-like actin-binding domain, respectively. In vitro assays revealed that MavH stimulates actin assembly on PI(3)P-containing liposomes causing their tubulation. In addition, the recruitment of CP by MavH negatively regulates F-actin density at the membrane. We further show that, in L. pneumophila-infected cells, MavH appears around the LCV at the very early stage of infection and facilitates bacterium entry into the host. Together, our results reveal a novel mechanism of membrane tubulation induced by membrane-dependent actin polymerization catalyzed by MavH that contributes to the early stage of L. pneumophila infection by regulating host actin dynamics.
56. Yin LM, Kudryashov DS, Zervas CG, Murk K.Front Cell Dev Biol. (2023) Nov 8;11:1329219. doi: 10.3389/fcell.2023.1329219. eCollection 2023. PMID: 38020892
Editorial: Evolution, emerging functions and structure of actin-binding proteins, Volume II.
55. Rajan S, Kudryashov DS, Reisler E. Biomolecules. (2023) Feb 28;13(3):450. doi: 10.3390/biom13030450.
Actin Bundles Dynamics and Architecture.
Cells use the actin cytoskeleton for many of their functions, including their division, adhesion, mechanosensing, endo- and phagocytosis, migration, and invasion. Actin bundles are the main constituent of actin-rich structures involved in these processes. An ever-increasing number of proteins that crosslink actin into bundles or regulate their morphology is being identified in cells. With recent advances in high-resolution microscopy and imaging techniques, the complex process of bundles formation and the multiple forms of physiological bundles are beginning to be better understood. Here, we review the physiochemical and biological properties of four families of highly conserved and abundant actin-bundling proteins, namely, α-actinin, fimbrin/plastin, fascin, and espin. We describe the similarities and differences between these proteins, their role in the formation of physiological actin bundles, and their properties-both related and unrelated to their bundling abilities. We also review some aspects of the general mechanism of actin bundles formation, which are known from the available information on the activity of the key actin partners involved in this process.
2022
54. Kudryashova E, Ankita, Ulrichs H, Shekhar S, Kudryashov DS. Science Advances. (2022) 8(46):eadc9239. doi: 10.1126/sciadv.adc9239.
Pointed-end processive elongation of actin filaments by Vibrio effectors VopF and VopL
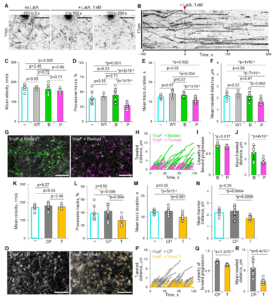 Abstract. According to the cellular actin dynamics paradigm, filaments grow at their barbed ends and depolymerize predominantly from their pointed ends to form polar structures and do productive work. We show that actin can elongate at the pointed end when assisted by Vibrio VopF/L toxins, which act as processive polymerases. In cells, processively moving VopF/L speckles are inhibited by factors blocking the pointed but not barbed ends. Multispectral single-molecule imaging confirmed that VopF molecules associate with the pointed end, actively promoting its elongation even in the presence of profilin. Consequently, VopF/L can break the actin cytoskeleton’s polarity by compromising actin-based cellular processes. Therefore, actin filament design allows processive growth at both ends, which suggests unforeseen possibilities for cellular actin organization, particularly in specialized cells and compartments.
Abstract. According to the cellular actin dynamics paradigm, filaments grow at their barbed ends and depolymerize predominantly from their pointed ends to form polar structures and do productive work. We show that actin can elongate at the pointed end when assisted by Vibrio VopF/L toxins, which act as processive polymerases. In cells, processively moving VopF/L speckles are inhibited by factors blocking the pointed but not barbed ends. Multispectral single-molecule imaging confirmed that VopF molecules associate with the pointed end, actively promoting its elongation even in the presence of profilin. Consequently, VopF/L can break the actin cytoskeleton’s polarity by compromising actin-based cellular processes. Therefore, actin filament design allows processive growth at both ends, which suggests unforeseen possibilities for cellular actin organization, particularly in specialized cells and compartments.
This discovery of thought-to-be-impossible processive polymerization at the actin pointed end in vertebrate cells was highlighted by 16 News Outlets
53. Dong S, Zheng W, Pinkerton N, Hansen J, Tikunova SB, Davis JP, Heissler SM, Kudryashova E, Egelman EH, Kudryashov DS. Int J Mol Sci. (2022) 23(13):7026. doi: 10.3390/ijms23137026.
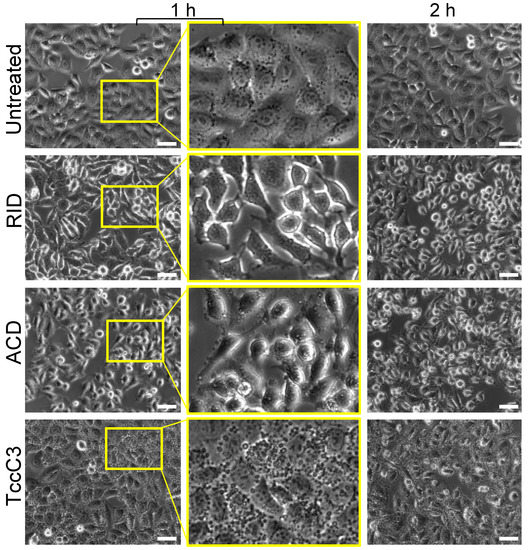
Photorhabdus luminescens TccC3 Toxin Targets the Dynamic Population of F-Actin and Impairs Cell Cortex Integrity
Abstract. Due to its essential role in cellular processes, actin is a common target for bacterial toxins. One such toxin, TccC3, is an effector domain of the ABC-toxin produced by entomopathogenic bacteria of Photorhabdus spp. Unlike other actin-targeting toxins, TccC3 uniquely ADP-ribosylates actin at Thr-148, resulting in the formation of actin aggregates and inhibition of phagocytosis. It has been shown that the fully modified F-actin is resistant to depolymerization by cofilin and gelsolin, but their effects on partially modified actin were not explored. We found that only F-actin unprotected by tropomyosin is the physiological TccC3 substrate. Yet, ADP-ribosylated G-actin can be produced upon cofilin-accelerated F-actin depolymerization, which was only mildly inhibited in partially modified actin. The affinity of TccC3-ADP-ribosylated G-actin for profilin and thymosin-β4 was weakened moderately but sufficiently to potentiate spontaneous polymerization in their presence. Interestingly, the Arp2/3-mediated nucleation was also potentiated by T148-ADP-ribosylation. Notably, even partially modified actin showed reduced bundling by plastins and α-actinin. In agreement with the role of these and other tandem calponin-homology domain actin organizers in the assembly of the cortical actin network, TccC3 induced intense membrane blebbing in cultured cells. Overall, our data suggest that TccC3 imposes a complex action on the cytoskeleton by affecting F-actin nucleation, recycling, and interaction with actin-binding proteins involved in the integration of actin filaments with each other and cellular elements.
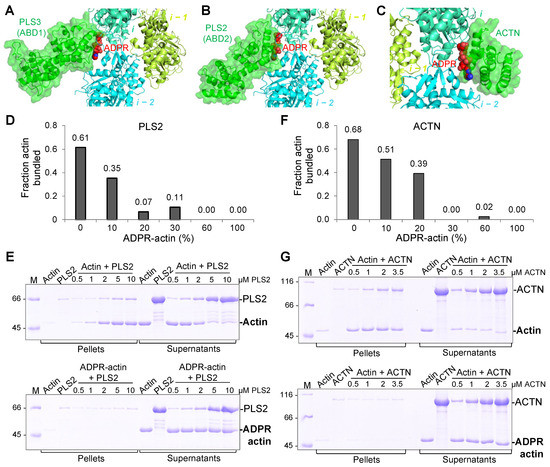
52. Schwebach CL, Kudryashova E, Agrawal R, Zheng W, Egelman EH, Kudryashov DS. Allosteric regulation controls actin-bundling properties of human plastins.
Nat Struct Mol Biol. 2022 Jun;29(6):519-528. doi: 10.1038/s41594-022-00771-1
Abstract: Plastins/fimbrins are conserved actin-bundling proteins contributing to motility, cytokinesis and other cellular processes by organizing strikingly different actin assemblies as in aligned bundles and branched networks. We propose that this ability of human plastins stems from an allosteric communication between their actin-binding domains (ABD1/2) engaged in a tight spatial association. Here we show that ABD2 can bind actin three orders of magnitude stronger than ABD1, unless the domains are involved in an equally strong inhibitory engagement. A mutation mimicking physiologically relevant phosphorylation at the ABD1-ABD2 interface greatly weakened their association, dramatically potentiating actin cross-linking. Cryo-EM reconstruction revealed the ABD1-actin interface and enabled modeling of the plastin bridge and domain separation in parallel bundles. We predict that a strong and tunable allosteric inhibition between the domains allows plastins to modulate the cross-linking strength, contributing to remodeling of actin assemblies of different morphologies defining the unique place of plastins in actin organization.

51. Kraus, J, Russell RW, Kudryashova E, Xu C, Katayala N, Perilla JR, Kudryashov DS, Polenova T (2022)
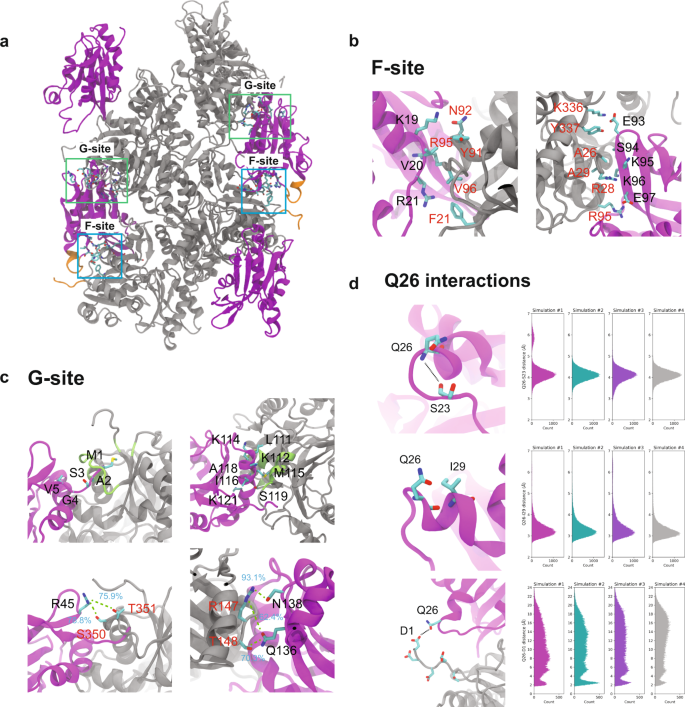
Magic angle spinning NMR structure of human cofilin-2 assembled on actin filaments reveals isoform-specific conformation and binding mode.
Nature communications 13(1):2114 doi:10.138/s41467-022029595-9
Abstract: Actin polymerization dynamics regulated by actin-binding proteins are essential for various cellular functions. The cofilin family of proteins are potent regulators of actin severing and filament disassembly. The structural basis for cofilin-isoform-specific severing activity is poorly understood as their high-resolution structures in complex with filamentous actin (F-actin) are lacking. Here, we present the atomic-resolution structure of the muscle-tissue-specific isoform, cofilin-2 (CFL2), assembled on ADP-F-actin, determined by magic-angle-spinning (MAS) NMR spectroscopy and data-guided molecular dynamics (MD) simulations. We observe an isoform-specific conformation for CFL2. This conformation is the result of a unique network of hydrogen bonding interactions within the α2 helix containing the non-conserved residue, Q26. Our results indicate F-site interactions that are specific between CFL2 and ADP-F-actin, revealing mechanistic insights into isoform-dependent F-actin disassembly.
2021
Find updates on other 2021 lab activities here!
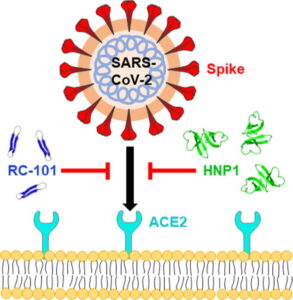
50. Elena Kudryashova, Ashley Zani, Geraldine Vilmen, Amit Sharma, Wuyuan Lu, Jacob S. Yount, Dmitri S. Kudryashov (2021). Inhibition of SARS-CoV-2 Infection by Human Defensin HNP1 and Retrocyclin RC-101. Journal of Molecular Biology https://doi.org/10.1016/j.jmb.2021.167225
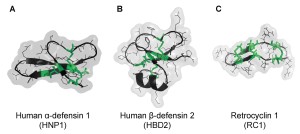
Abstract: Severe acute respiratory syndrome coronavirus (SARS-CoV)-2 is an enveloped virus responsible for the COVID-19 pandemic. The emergence of new potentially more transmissible and vaccine-resistant variants of SARS-CoV-2 is an ever-present threat. Thus, it remains essential to better understand innate immune mechanisms that can inhibit the virus. One component of the innate immune system with broad antipathogen, including antiviral, activity is a group of cationic immune peptides termed defensins. The ability of defensins to neutralize enveloped and non-enveloped viruses and to inactivate numerous bacterial toxins correlate with their ability to promote the unfolding of proteins with high conformational plasticity. We found that human neutrophil α-defensin HNP1 binds to SARS-CoV-2 Spike protein with submicromolar affinity that is more than 20 fold stronger than its binding to serum albumin. As such, HNP1, as well as a θ-defensin retrocyclin RC-101, both interfere with Spike-mediated membrane fusion, Spike-pseudotyped lentivirus infection, and authentic SARS-CoV-2 infection in cell culture. These effects correlate with the abilities of the defensins to destabilize and precipitate Spike protein and inhibit the interaction of Spike with the ACE2 receptor. Serum reduces the anti-SARS-CoV-2 activity of HNP1, though at high concentrations, HNP1 was able to inactivate the virus even in the presence of serum. Overall, our results suggest that defensins can negatively affect the native conformation of SARS-CoV-2 Spike, and that α- and θ-defensins may be valuable tools in developing SARS-CoV-2 infection prevention strategies.
Keywords: SARS-CoV-2; innate immunity; defensins; human neutrophil peptide HNP1; retrocyclin RC-101
49. Schwebach, C. L., Kudryashova, E., & Kudryashov, D. S. (2021). Plastin 3 in X-Linked Osteoporosis: Imbalance of Ca2+-Dependent Regulation Is Equivalent to Protein Loss. Frontiers in Cell and Developmental Biology, 8(January), 1–8. https://doi.org/10.3389/fcell.2020.635783
Abstract: Osteogenesis imperfecta is a genetic disorder disrupting bone development and remodeling. The primary causes of osteogenesis imperfecta are pathogenic variants of collagen and collagen processing genes. However, recently variants of the actin bundling protein plastin 3 have been identified as another source of osteogenesis imperfecta. Plastin 3 is a highly conserved protein involved in several important cellular structures and processes and is controlled by intracellular Ca2+ which potently inhibits its actin-bundling activity. The precise mechanisms by which plastin 3 causes osteogenesis imperfecta remain unclear, but recent advances have contributed to our understanding of bone development and the actin cytoskeleton. Here, we review the link between plastin 3 and osteogenesis imperfecta highlighting in vitro studies and emphasizing the importance of Ca2+ regulation in the localization and functionality of plastin 3.
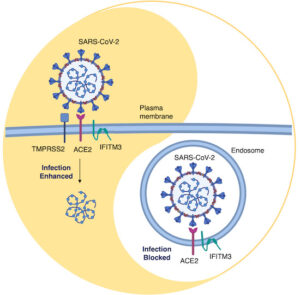
48. Shi, G., Kenney, A. D., Kudryashova, E., Zani, A., Zhang, L., Lai, K. K., … Yount, J. S. (2021). Opposing activities of IFITM proteins in SARS‐CoV‐2 infection. The EMBO Journal, 40(3), 1–12. https://doi.org/10.15252/embj.2020106501
Abstract: Interferon‐induced transmembrane proteins (IFITMs) restrict infections by many viruses, but a subset of IFITMs enhance infections by specific coronaviruses through currently unknown mechanisms. We show that SARS‐CoV‐2 Spike‐pseudotyped virus and genuine SARS‐CoV‐2 infections are generally restricted by human and mouse IFITM1, IFITM2, and IFITM3, using gain‐ and loss‐of‐function approaches. Mechanistically, SARS‐CoV‐2 restriction occurred independently of IFITM3 S‐palmitoylation, indicating a restrictive capacity distinct from reported inhibition of other viruses. In contrast, the IFITM3 amphipathic helix and its amphipathic properties were required for virus restriction. Mutation of residues within the IFITM3 endocytosis‐promoting YxxФ motif converted human IFITM3 into an enhancer of SARS‐CoV‐2 infection, and cell‐to‐cell fusion assays confirmed the ability of endocytic mutants to enhance Spike‐mediated fusion with the plasma membrane. Overexpression of TMPRSS2, which increases plasma membrane fusion versus endosome fusion of SARS‐CoV‐2, attenuated IFITM3 restriction and converted amphipathic helix mutants into infection enhancers. In sum, we uncover new pro‐ and anti‐viral mechanisms of IFITM3, with clear distinctions drawn between enhancement of viral infection at the plasma membrane and amphipathicity‐based mechanisms used for endosomal SARS‐CoV‐2 restriction.
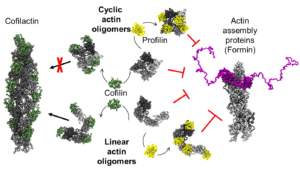
47. Smith, H., Pinkerton, N., Heisler, D. B., Kudryashova, E., Hall, A. R., Karch, K. R., … Kudryashov, D. S. (2021). Rounding out the understanding of ACD toxicity with the discovery of cyclic forms of actin oligomers. International Journal of Molecular Sciences, 22(2), 1–19. https://doi.org/10.3390/ijms22020718
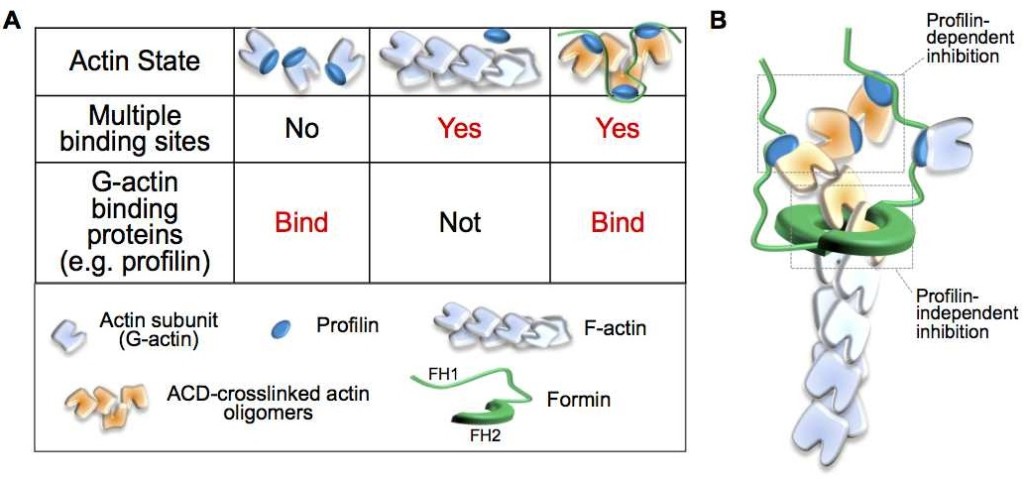
Abstract: Actin is an essential element of both innate and adaptive immune systems and can aid in motility and translocation of bacterial pathogens, making it an attractive target for bacterial toxins. Pathogenic Vibrio and Aeromonas genera deliver actin cross-linking domain (ACD) toxin into the cytoplasm of the host cell to poison actin regulation and promptly induce cell rounding. At early stages of toxicity, ACD covalently cross-links actin monomers into oligomers (AOs) that bind through multivalent interactions and potently inhibit several families of actin assembly proteins. At advanced toxicity stages, we found that the terminal protomers of linear AOs can get linked together by ACD to produce cyclic AOs. When tested against formins and Ena/VASP, linear and cyclic AOs exhibit similar inhibitory potential, which for the cyclic AOs is reduced in the presence of profilin. In coarse-grained molecular dynamics simulations, profilin and WH2-motif binding sites on actin subunits remain exposed in modeled AOs of both geometries. We speculate, therefore, that the reduced toxicity of cyclic AOs is due to their reduced configurational entropy. A characteristic feature of cyclic AOs is that, in contrast to the linear forms, they cannot be straightened to form filaments (e.g., through stabilization by cofilin), which makes them less susceptible to neutralization by the host cell.
2020
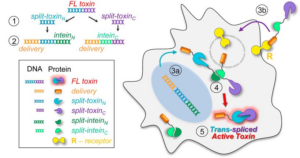
46. Purde, V., Kudryashova, E., Heisler, D. B., Shakya, R., & Kudryashov, D. S. (2020). Intein-mediated cytoplasmic reconstitution of a split toxin enables selective cell ablation in mixed populations and tumor xenografts. Proceedings of the National Academy of Sciences of the United States of America, 117(36), 22090–22100. https://doi.org/10.1073/pnas.2006603117
Abstract: The application of proteinaceous toxins for cell ablation is limited by their high on- and off-target toxicity, severe side effects, and a narrow therapeutic window. The selectivity of targeting can be improved by intein-based toxin reconstitution from two dysfunctional fragments provided their cytoplasmic delivery via independent, selective pathways. While the reconstitution of proteins from genetically encoded elements has been explored, exploiting cell-surface receptors for boosting selectivity has not been attained. We designed a robust splitting algorithm and achieved reliable cytoplasmic reconstitution of functional diphtheria toxin from engineered intein-flanked fragments upon receptor-mediated delivery of one of them to the cells expressing the counterpart. Retargeting the delivery machinery toward different receptors overexpressed in cancer cells enables selective ablation of specific subpopulations in mixed cell cultures. In a mouse model, the transmembrane delivery of a split-toxin construct potently inhibits the growth of xenograft tumors expressing the split counterpart. Receptor-mediated delivery of engineered split proteins provides a platform for precise therapeutic and experimental ablation of tumors or desired cell populations while also greatly expanding the applicability of the intein-based protein transsplicing.
This publication was highlighted in several media outlets:
-
-
-
Splitting immunotoxins could increase specificity toward cancers
-
In One Cancer Therapy, Two Halves Are Safer Than A Whole
-
Splitting immunotoxins in half could increase their specificity toward cancers, study suggests
-
Delivering split immunotoxins separately to cancer cells can reduce life-threatening side effects
-
Splitting Cancer Drug is Safer Than A Whole
-
-
45. Christopher L. Schwebach, Elena Kudryashova, Weili Zheng, Matthew Orchard, Harper Smith, Lucas Runyan, Edward Egelman, Dmitri Kudryashov (2020) Osteogenesis imperfecta mutations in plastin 3 lead to impaired calcium regulation of actin bundling. Bone Research 8-21 (2020). https://rdcu.be/b4lcy
Abstract: Mutations in actin-bundling protein plastin 3 (PLS3) emerged as a cause of congenital osteoporosis, but neither the role of PLS3 in bone development nor the mechanisms underlying PLS3-dependent osteoporosis are understood. Of the over 20 identified osteoporosis-linked PLS3 mutations, we investigated all five that are expected to produce full-length protein. One of the mutations distorted an actin-binding loop in the second actin-binding domain of PLS3 and abolished F-actin bundling as revealed by cryo-EM reconstruction and protein interaction assays. Surprisingly, the remaining four mutants fully retained F-actin bundling ability. However, they displayed defects in Ca²⁺ sensitivity: two of the mutants lost the ability to be inhibited by Ca²⁺, while the other two became hypersensitive to Ca²⁺. Each group of the mutants with similar biochemical properties showed highly characteristic cellular behavior. Wild-type PLS3 was distributed between lamellipodia and focal adhesions. In striking contrast, the Ca²⁺-hyposensitive mutants were not found at the leading edge but localized exclusively at focal adhesions/stress fibers, which displayed reinforced morphology. Consistently, the Ca²⁺-hypersensitive PLS3 mutants were restricted to lamellipodia, while chelation of Ca²⁺ caused their redistribution to focal adhesions. Finally, the bundling-deficient mutant failed to co-localize with any F-actin structures in cells despite a preserved F-actin binding through a non-mutation-bearing actin-binding domain. Our findings revealed that severe osteoporosis can be caused by a mutational disruption of the Ca²⁺-controlled PLS3’s cycling between adhesion complexes and the leading edge. Integration of the structural, biochemical, and cell biology insights enabled us to propose a molecular mechanism of plastin activity regulation by Ca²⁺.
2019
44. Purde V, Busch F, Kudryashova E, Wysocki VH, Kudryashov DS. (2019) Oligomerization Affects the Ability of Human Cyclase-Associated Proteins 1 and 2 to Promote Actin Severing by Cofilins. International Journal of Molecular Sciences 20(22):5647. doi:10.3390/ijms20225647.
Abstract: Actin-depolymerizing factor (ADF)/cofilins accelerate actin turnover by severing aged actin filaments and promoting the dissociation of actin subunits. In the cell, ADF/cofilins are assisted by other proteins, among which cyclase-associated proteins 1 and 2 (CAP1,2) are particularly important. The N-terminal half of CAP has been shown to promote actin filament dynamics by enhancing ADF-/cofilin-mediated actin severing, while the central and C-terminal domains are involved in recharging the depolymerized ADP-G-actin/cofilin complexes with ATP and profilin. We analyzed the ability of the N-terminal fragments of human CAP1 and CAP2 to assist human isoforms of "muscle" (CFL2) and "non-muscle" (CFL1) cofilins in accelerating actin dynamics. By conducting bulk actin depolymerization assays and monitoring single-filament severing by total internal reflection fluorescence (TIRF) microscopy, we found that the N-terminal domains of both isoforms enhanced cofilin-mediated severing and depolymerization at similar rates. According to our analytical sedimentation and native mass spectrometry data, the N-terminal recombinant fragments of both human CAP isoforms form tetramers. Replacement of the original oligomerization domain of CAPs with artificial coiled-coil sequences of known oligomerization patterns showed that the activity of the proteins is directly proportional to the stoichiometry of their oligomerization; i.e., tetramers and trimers are more potent than dimers, which are more effective than monomers. Along with higher binding affinities of the higher-order oligomers to actin, this observation suggests that the mechanism of actin severing and depolymerization involves simultaneous or consequent and coordinated binding of more than one N-CAP domain to F-actin/cofilin complexes.
2018
43. Kudryashova E, Heisler DB, Williams B, Harker A, Shafer K, Quinlan ME, Kovar DR, Vavylonis D, Kudryashov DS (2018) Actin-cross-linking Toxin is a Universal Inhibitor of Tandem-organized and Oligomeric G-actin-binding Proteins. Current Biology In press.
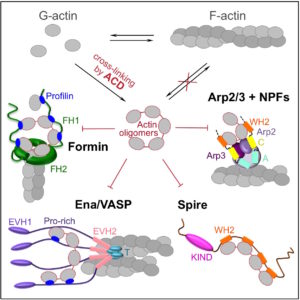
Abstract: Delivery of bacterial toxins to host cells is hindered by host protective barriers. This obstruction dictates a remarkable efficiency of toxins, a single copy of which may kill a host cell. Efficiency of actin-targeting toxins is further hampered by an overwhelming abundance of their target. The actin cross-linking domain (ACD) toxins of Vibrio species and related bacterial genera catalyze the formation of covalently cross-linked actin oligomers. Recently, we reported that the ACD toxicity can be amplified via a multivalent inhibitory association of actin oligomers with actin assembly factors formins, suggesting that the oligomers may act as secondary toxins. Importantly, many proteins involved in nucleation, elongation, severing, branching, and bundling of actin filaments contain G-actin-binding Wiskott-Aldrich syndrome protein (WASP)-homology motifs 2 (WH2) organized in tandem and therefore may act as a multivalent platform for high-affinity interaction with the ACD-cross-linked actin oligomers. Using live-cell single-molecule speckle (SiMS) microscopy, total internal reflection fluorescence (TIRF) microscopy, and actin polymerization assays, we show that, in addition to formins, the oligomers bind with high affinity and potently inhibit several families of actin assembly factors: Ena/vasodilator-stimulated phosphorprotein (VASP); Spire; and the Arp2/3 complex, both in vitro and in live cells. As a result, ACD blocks the actin retrograde flow and membrane dynamics and disrupts association of Ena/VASP with adhesion complexes. This study defines ACD as a universal inhibitor of tandem-organized G-actin binding proteins that overcomes the abundance of actin by redirecting the toxicity cascade toward less abundant targets and thus leading to profound disorganization of the actin cytoskeleton and disruption of actin-dependent cellular functions.
This work has been highlighted in:
2017
42. Schwebach CL, Agrawal R; Lindert S, Kudryashova E, Kudryashov DS (2017) The roles of Actin-Binding Domains 1 and 2 in the Calcium-dependent Regulation of Actin Filament Bundling by Human Plastins. Journal of Molecular Biology 429(16):2490-2508 https://doi.org/10.1016/j.jmb.2017.06.021.
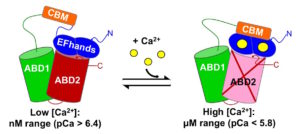
Abstract: The actin cytoskeleton is a complex network controlled by a vast array of intricately regulated actin-binding proteins. Human plastins (PLS1, PLS2, and PLS3) are evolutionary conserved proteins that non-covalently crosslink actin filaments into tight bundles. Through stabilization of such bundles, plastins contribute, in an isoform-specific manner, to the formation of kidney and intestinal microvilli, inner ear stereocilia, immune synapses, endocytic patches, adhesion contacts, and invadosomes of immune and cancer cells. All plastins comprise an N-terminal Ca2+-binding regulatory headpiece domain followed by two actin-binding domains (ABD1 and ABD2). Actin bundling occurs due to simultaneous binding of both ABDs to separate actin filaments. Bundling is negatively regulated by Ca2+, but the mechanism of this inhibition remains unknown. In this study, we found that the bundling abilities of PLS1 and PLS2 were similarly sensitive to Ca2+ (pCa50 ~6.4), whereas PLS3 was less sensitive (pCa50 ~5.9). At the same time, all three isoforms bound to F-actin in a Ca2+-independent manner, suggesting that binding of only one of the ABDs is inhibited by Ca2+. Using limited proteolysis and mass spectrometry, we found that in the presence of Ca2+ the EF-hands of human plastins bound to an immediately adjacent sequence homologous to canonical calmodulin-binding peptides. Furthermore, our data from differential centrifugation, Förster resonance energy transfer, native electrophoresis, and chemical crosslinking suggest that Ca2+ does not affect ABD1 but inhibits the ability of ABD2 to interact with actin. A structural mechanism of signal transmission from Ca2+ to ABD2 through EF-hands remains to be established.
41. Kudryashova E, Seveau S, Kudryashov DS (2017) Invited review – Targeting and inactivation of bacterial toxins by human defensins. Biological Chemistry ISSN (Online) 1437-4315, ISSN (Print) 1431-6730, DOI: https://doi.org/10.1515/hsz-2017-0106.
Abstract: Defensins, as a prominent family of antimicrobial peptides (AMP), are major effectors of the innate immunity with a broad range of immune modulatory and antimicrobial activities. In particular, defensins are the only recognized fast-response molecules that can neutralize a broad range of bacterial toxins, many of which are among the deadliest compounds on the planet. For a decade, the mystery of how a small and structurally conserved group of peptides can neutralize a heterogeneous group of toxins with little to no sequential and structural similarity remained unresolved. Recently, it was found that defensins recognize and target structural plasticity/thermodynamic instability, fundamental physicochemical properties that unite many bacterial toxins and distinguish them from the majority of host proteins. Binding of human defensins promotes local unfolding of the affected toxins, destabilizes their secondary and tertiary structures, increases susceptibility to proteolysis, and leads to their precipitation. While the details of toxin destabilization by defensins remain obscure, here we briefly review properties and activities of bacterial toxins known to be affected by or resilient to defensins, and discuss how recognized features of defensins correlate with the observed inactivation.
40. Harterink M, da Silva ME, Will L, Turan J, Ibrahim A, Lang AE, van Battum EY, Pasterkamp RJ, Kapitein LC, Kudryashov DS, Barres BA, Hoogenraad CC, Zuchero JB (2017) DeActs: genetically encoded tools for perturbing the actin cytoskeleton in single cells. Nature Methods doi: 10.1038/nmeth.4257
Abstract: The actin cytoskeleton is essential for many fundamental biological processes, but tools for directly manipulating actin dynamics are limited to cell-permeable drugs that preclude single-cell perturbations. Here we describe DeActs, genetically encoded actin-modifying polypeptides, which effectively induce actin disassembly in eukaryotic cells. We demonstrate that DeActs are universal tools for studying the actin cytoskeleton in single cells in culture, tissues, and multicellular organisms including various neurodevelopmental model systems.
39. Yehl J, Kudryashova E, Reisler E, Kudryashov DS, Polenova T (2017) Structural Analysis of Human Cofilin 2/Filamentous Actin Assemblies: Atomic-Resolution Insights from Magic Angle Spinning NMR Spectroscopy. Scientific Reports 7, 44506 doi:10.1038/srep44506
Abstract: Cellular actin dynamics is an essential element of numerous cellular processes, such as cell motility, cell division and endocytosis. Actin’s involvement in these processes is mediated by many actin-binding proteins, among which the cofilin family plays unique and essential role in accelerating actin treadmilling in filamentous actin (F-actin) in a nucleotide-state dependent manner. Cofilin preferentially interacts with older filaments by recognizing time-dependent changes in F-actin structure associated with the hydrolysis of ATP and release of inorganic phosphate (Pi) from the nucleotide cleft of actin. The structure of cofilin on F-actin and the details of the intermolecular interface remain poorly understood at atomic resolution. Here we report atomic-level characterization by magic angle spinning (MAS) NMR of the muscle isoform of human cofilin 2 (CFL2) bound to F-actin. We demonstrate that resonance assignments for the majority of atoms are readily accomplished and we derive the intermolecular interface between CFL2 and F-actin. The MAS NMR approach reported here establishes the foundation for atomic-resolution characterization of a broad range of actin-associated proteins bound to F-actin.
38. Kudryashova E, Heisler DB, Kudryashov DS. (2017) Book chapter “Pathogenic  mechanisms of actin cross-linking toxins – peeling away the layers” in Spinger book series “Current Topics in Microbiology and Immunology” (CTMI) 399:87-112 “Actin Cytoskeleton and Bacterial Infection”. Mannherz HG (Ed.) ISBN 978-3-319-50047-8
mechanisms of actin cross-linking toxins – peeling away the layers” in Spinger book series “Current Topics in Microbiology and Immunology” (CTMI) 399:87-112 “Actin Cytoskeleton and Bacterial Infection”. Mannherz HG (Ed.) ISBN 978-3-319-50047-8
Abstract: Actin cross-linking toxins are produced by Gram-negative bacteria from Vibrio and Aeromonas genera. The toxins were named actin cross-linking domains (ACD), since the first and most of the subsequently discovered ACDs were found as effector domains in larger MARTX and VgrG toxins. Among recognized human pathogens, ACD is produced by Vibrio cholerae, Vibrio vulnificus, and Aeromonas hydrophila. Upon delivery to the cytoplasm of a host cell, ACD covalently cross-links actin monomers into non-polymerizable actin oligomers of various lengths. Provided sufficient doses of toxin are delivered, most or all actin can be promptly cross-linked into non-functional oligomers, leading to cell rounding, detachment from the substrate and, in many cases, cell death. Recently, a deeper layer of ACD toxicity with a less obvious but more potent mechanism was discovered. According to this finding, low doses of the ACD-produced actin oligomers can actively disrupt the actin cytoskeleton by potently inhibiting essential actin assembly proteins, formins. The first layer of toxicity is direct (as actin is the immediate and the only target), passive (since ACD-cross-linked actin oligomers are toxic only because they are non-functional), and less potent (as bulk quantities of one of the most abundant cytoplasmic proteins, actin, have to be modified). The second mechanism is indirect (as major targets, formins, are not affected by ACD directly), active (because actin oligomers act as "secondary" toxins), and highly potent [as it affects scarce and essential actin-binding proteins (ABPs)].
2016
37. Kudryashova E, Koneru PC, Kvaratskhelia M, Strömstedt AA, Lu W, Kudryashov DS (2016) Thermodynamic instability of viral proteins is a pathogen-associated molecular pattern targeted by human defensins. Scientific Reports 6, 32499 doi:10.1038/srep32499
Abstract: Human defensins are innate immune defense peptides with a remarkably broad repertoire of anti-pathogen activities. In addition to modulating immune response, inflammation, and angiogenesis, disintegrating bacterial membranes, and inactivating bacterial toxins, defensins are known to intercept various viruses at different stages of their life cycles, while remaining relatively benign towards human cells and proteins. Recently we have found that human defensins inactivate proteinaceous bacterial toxins by taking advantage of their low thermodynamic stability and acting as natural “anti-chaperones”, i.e. destabilizing the native conformation of the toxins. In the present study we tested various proteins produced by several viruses (HIV-1, PFV, and TEV) and found them to be susceptible to destabilizing effects of human α-defensins HNP-1 and HD-5 and the synthetic θ-defensin RC-101, but not β-defensins hBD-1 and hBD-2 or structurally related plant-derived peptides. Defensin-induced unfolding promoted exposure of hydrophobic groups otherwise confined to the core of the viral proteins. This resulted in precipitation, an enhanced susceptibility to proteolytic cleavage, and a loss of viral protein activities. We propose, that defensins recognize and target a common and essential physico-chemical property shared by many bacterial toxins and viral proteins – the intrinsically low thermodynamic protein stability.
36. Serebryannyy LA, Parilla M, Annibale P, Cruz CM, Laster K, Gratton E, Kudryashov D, Kosak ST, Gottardi CJ, de Lanerolle P. (2016) Persistent nuclear actin filaments inhibit transcription by RNA polymerase II. Journal of Cell Science 2016 Sep 15;129(18):3412-25. doi: 10.1242/jcs.195867
2015
35. Vilitkevich EL, Khapchaev AY, Kudryashov DS, Nikashin AV, Schavocky JP, Lukas TJ, Watterson DM, Shirinsky VP (2015) Phosphorylation Regulates Interaction of 210-kDa Myosin Light Chain Kinase N-terminal Domain with Actin Cytoskeleton. Biochemistry (Mosc). 80(10):1288-97 doi:10.1134/S0006297915100090
34. Kudryashova E, Lu W, Kudryashov DS (2015) “Defensins versus pathogens: an unfolding story” – Editorial. Oncotarget 6(30):28533-28534 doi:10.18632/oncotarget.5109
33. Heisler DB, Kudryashova E, Grinevich DO, Suarez C, Winkelman JD, Birukov KG, Kotha SR, Parinandi NL, Vavylonis D, Kovar DR, Kudryashov DS (2015) “ACD toxin–produced actin oligomers poison formin-controlled actin polymerization.” Science 349(6247):535-539 doi: 10.1126/science.aab4090
Reprint: http://www.sciencemag.org/cgi/rapidpdf/349/6247/535?ijkey=F1yNiUEUjtzME&keytype=ref&siteid=sci
Full Text: http://www.sciencemag.org/cgi/content/full/349/6247/535?ijkey=F1yNiUEUjtzME&keytype=ref&siteid=sci
Abstract: The actin cross-linking domain (ACD) is an actin-specific toxin produced by several pathogens, including life-threatening spp. of Vibrio cholerae, Vibrio vulnificus, and Aeromonas hydrophila. Actin cross-linking by ACD is thought to lead to slow cytoskeleton failure owing to a gradual sequestration of actin in the form of nonfunctional oligomers. Here, we found that ACD converted cytoplasmic actin into highly toxic oligomers that potently “poisoned” the ability of major actin assembly proteins, formins, to sustain actin polymerization. Thus, ACD can target the most abundant cellular protein by using actin oligomers as secondary toxins to efficiently subvert cellular functions of actin while functioning at very low doses.
EDITOR'S SUMMURY: A little toxin can do a lot. The actin cross-linking domain (ACD) is an actin-specific toxin produced by several bacterial pathogens. Heisler et al. discovered that ACD's pathogenic mechanism involves a highly unusual toxicity amplification cascade. Rather than directly inactivating the actin cytoskeleton, ACD blocks the activity of formins, actin regulatory proteins that play crucial roles in numerous cellular activities. ACD is exceptionally potent, even though its substrate is the most abundant protein of a eukaryotic cell: actin.
This work has been highlighted in:
-
-
- Ohio State Research NEWS RELEASE
- Genetic Engineering & Biotechnology News (GEN)
- CHEMIE.DE information portal Bionity.com
- NATURE REVIEWS MICROBIOLOGY | RESEARCH HIGHLIGHT – Nature Reviews Microbiology 13, 526 doi:10.1038/nrmicro3539
- EurekAlert! AAAS
- News Medical
- Microbe World
- Science Daily
- Science Newsline Biology
- Medical News Today
- NewsUnited
- NewsWise
- sott.net
- phys.org
-
32. Kudryashova E, Seveau S, Lu W, Kudryashov DS (2015) Retrocyclins neutralize bacterial toxins by potentiating their unfolding. Biochemical Journal 467:311–320 doi:10.1042/BJ20150049
Abstract: We demonstrate that retrocyclins, promising therapeutic peptides, neutralize bacterial toxins by inducing their unfolding and exposing hydrophobic regions, normally buried in the molecule interior, to solvents. Retrocyclin-induced toxin unfolding leads to toxin precipitation, proneness to proteolytic degradation, and abrogated activity.
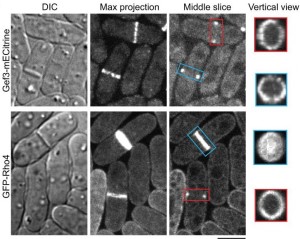
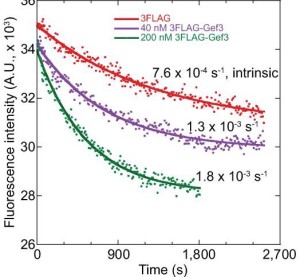
31. Wang N, Wang M, Zhu YH, Grosel TW, Sun D, Kudryashov DS, Wu JQ (2015) aaaThe Rho-GEF Gef3 interacts with the septin complex and activates the GTPase Rho4 during fission yeast cytokinesis. Molecular Biology of the Cell 26(2):238-255 doi:10.1091/mbc.E14-07-1196
A
A
A
Abstract: Rho GTPases, activated by Rho guanine nucleotide exchange factors (GEFs), are conserved molecular switches for signal transductions that regulate diverse cellular processes including cell polarization and cytokinesis. The fission yeast Schizosaccharomyces pombe has six Rho GTPases (Cdc42 and Rho1-Rho5) and seven Rho GEFs (Scd1, Rgf1-Rgf3, and Gef1-Gef3). The GEFs for Rho2-Rho5 have not been unequivocally assigned. Particularly, Gef3, the smallest Rho GEF, was barely studied. Here we show that Gef3 colocalizes with septins at the cell equator. Gef3 physically interacts with septins and anillin Mid2 and depends on them to localize. Gef3 co-precipitates with GDP-bound Rho4 in vitro and accelerates nucleotide exchange of Rho4, suggesting that Gef3 is a GEF for Rho4. Consistently, Gef3 and Rho4 are in the same genetic pathways to regulate septum formation and/or cell separation. In gef3Δ cells, the localizations of two potential Rho4 effectors, glucanases Eng1 and Agn1, are abnormal; and active Rho4 level is reduced, indicating that Gef3 is involved in Rho4 activation in vivo. Moreover, overexpression of active Rho4 or Eng1 rescues the septation defects of mutants containing gef3Δ. Together, our data support that Gef3 interacts with the septin complex and activates Rho4 GTPase as a Rho GEF for septation in fission yeast.
2014
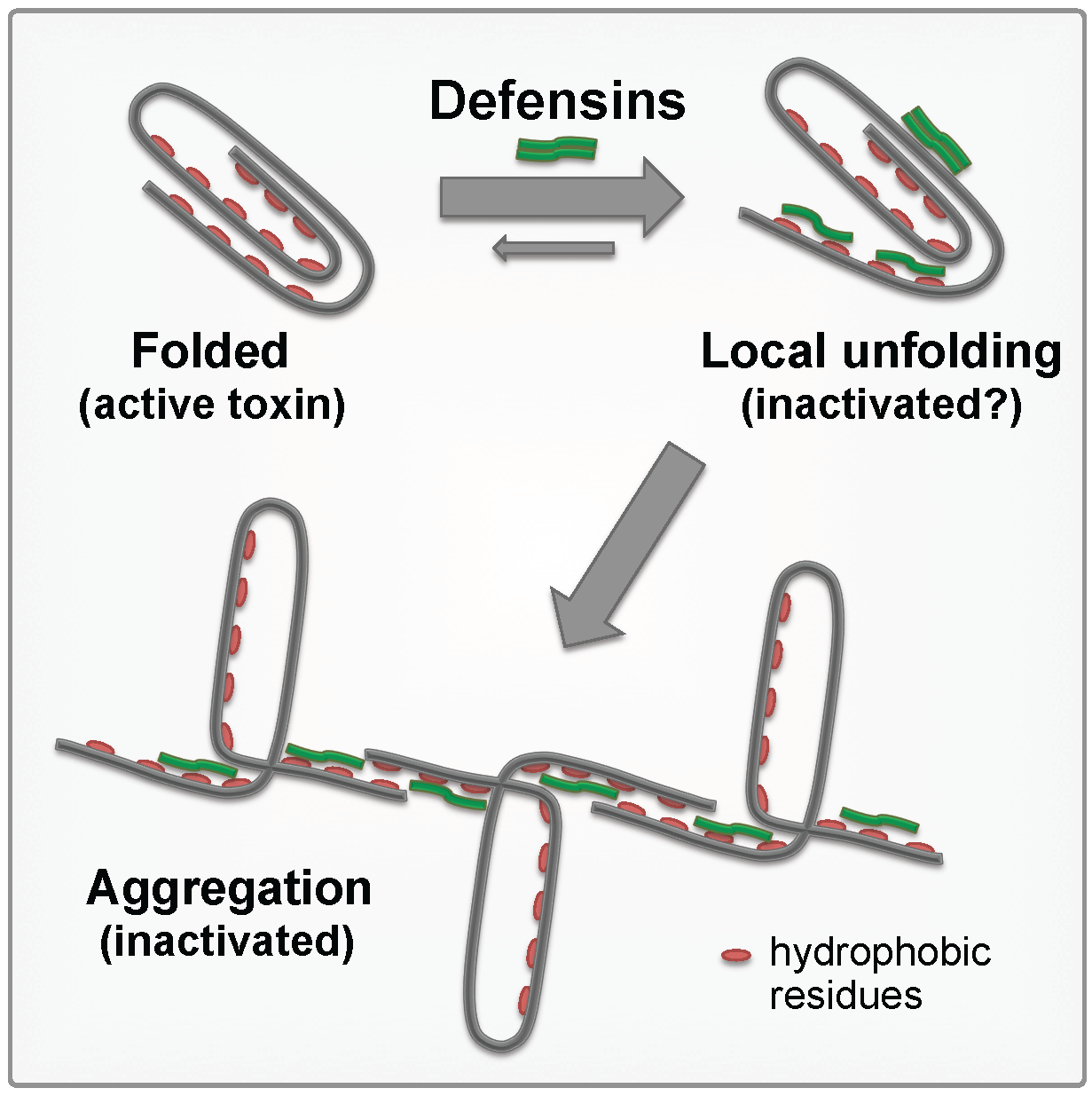
30. Kudryashova E., Quintyn R., Seveau S., Lu W., Wysocki V.H., Kudryashov D.S. (2014) aaaHuman defensins facilitate local unfolding of thermodynamically unstable regions of aaabacterial protein toxins. Immunity 41(5):709-721 doi:10.1016/j.immuni.2014.10.018
![]()
A
![]()
![]()
Abstract: Defensins are amphiphilic immune peptides capable of inactivating several classes of bacterial exotoxins by poorly understood mechanisms. We demonstrate that defensins take advantage of thermodynamic instability common for many bacterial toxins, cause their local unfolding, expose new regions for proteolysis, and potentiate precipitation through the exposure of hydrophobic interfaces.
This work has been highlighted in:
-
-
- Immunity 41(5), 2014 issue by Jens-Michael Shroder Revealing the Achilles Heel of Bacterial Toxins
- Recommended in F1000Prime as being of special significance in its field
- Ohio State Research NEWS RELEASE
- SciGuru SCIENCE NEWS “Research shows small proteins called defensins neutralize toxins released by pathogens”
- NSF SCIENCE360 NEWS (HEADLINES) “The best offense against bacteria is a good defense”
- OSU COLLEGE OF ARTS AND SCIENCES NEWS & UPDATES “New study shows small proteins that can disable bacterial toxins”
- NATURE IMMUNOLOGY | RESEARCH HIGHLIGHTS by Ioana Visan “Unfold and destroy” in Nature Immunology 16,141 (2015)
- Newsroom on NEWSWISE The Best Offense Against Bacteria is a Good Defense Research shows small proteins called defensins neutralize toxins released by pathogens Released: 7-Jan-2015 9:30 AM EST
- R&D Magazine
- Web Newswire
- BioPortfolio
- Science Daily
-
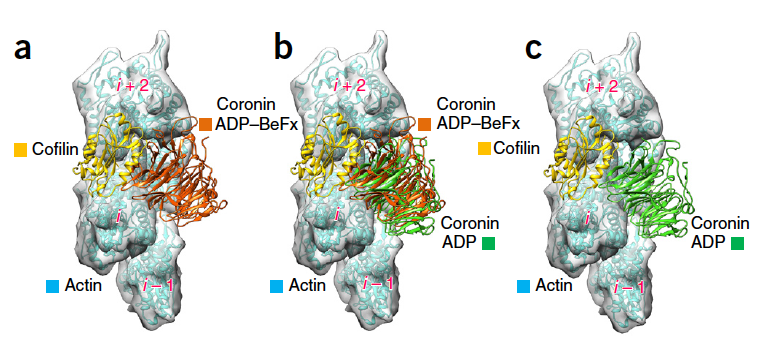
29. Ge P., Oztug Durer Z.A., Kudryashov D.S., Zhou Z.H., Reisler E. (2014) Cryo-EM reveals different coronin binding modes for ADP- and ADP-BeFx actin filaments. Nature Structural & Molecular Biology 21:1075–1081 doi: 10.1038/nsmb.2907
A
A
-
-
- Recommended in F1000Prime as being of special significance in its field by F1000 Faculty Member Bruce Goode [DOI: 10.3410/f.725223799.793505444]
-
Abstract: Essential cellular processes involving the actin cytoskeleton are regulated by auxiliary proteins that can sense the nucleotide state of actin. Here we report cryo-EM structures for ADP-bound and ADP–beryllium fluoride (ADP–BeFx, an ADP-Pi mimic)-bound actin filaments in complex with the b-propeller domain of yeast coronin 1 (crn1), at 8.6-Å resolution.
28. Kudryashova E., Heisler D., Zywiec A., Kudryashov D.S. (2014) Thermodynamic aaaproperties of the effector domains of MARTX toxins suggest their unfolding for aaatranslocation across the host membrane. Molecular Microbiology 92(5):1056–1071 aaadoi: 10.1111/mmi.12615 pdf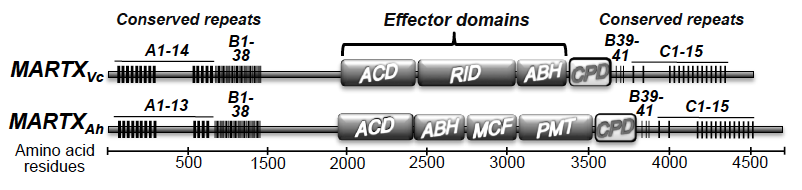
Abstract: MARTX (multifunctional autoprocessing repeats-in-toxin) family toxins are produced by Vibrio cholerae, Vibrio vulnificus, Aeromonas hydrophila and other Gram-negative bacteria. Effector domains of MARTX toxins cross the cytoplasmic membrane of a host cell through a putative pore formed by the toxin's glycine-rich repeats. The structure of the pore is unknown and the translocation mechanism of the effector domains is poorly understood. We found that all but one effector domains of V. cholerae and A. hydrophila MARTX toxins are thermodynamically unstable and several acquire a molten globule state near human physiological temperatures. This study extends the list of thermolabile bacterial toxins, suggesting that this quality is essential and could be susceptible for selective targeting of pathogenic toxins.
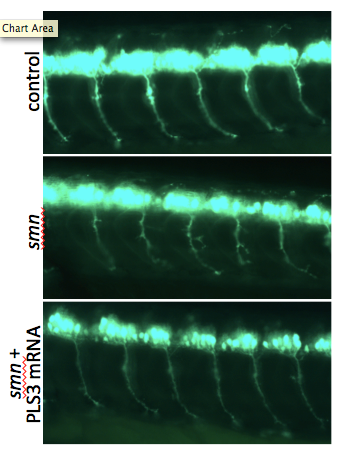
27. Lyon A.N., Pineda J.R.H., Kudryashova E., Kudryashov D.S., Beattie C.E. (2014) aaaCalcium binding is essential for plastin 3 function in Smn-deficient motor neurons. aaaHuman Molecular Genetics 23(8):1990-2004 doi: 10.1093/hmg/ddt595 pdf
 A
A 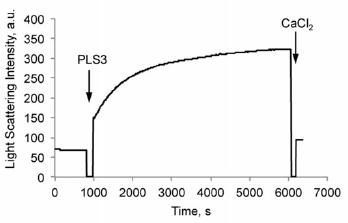 A
A ![]()
Abstract: The actin-binding and bundling protein, plastin 3 (PLS3), was identified as a protective modifier of spinal muscular atrophy (SMA) in some patient populations and as a disease modifier in animal models of SMA. To determine how PLS3 functions in SMA, we generated deletion constructs of conserved PLS3 structural domains. Our results indicate that Ca(2+) regulation is essential for the function of PLS3 in motor axons.
2013
26. Kudryashov D.S., Reisler E. (2013) ATP and ADP actin states. Biopolymers aaa99(4):245–256 pdf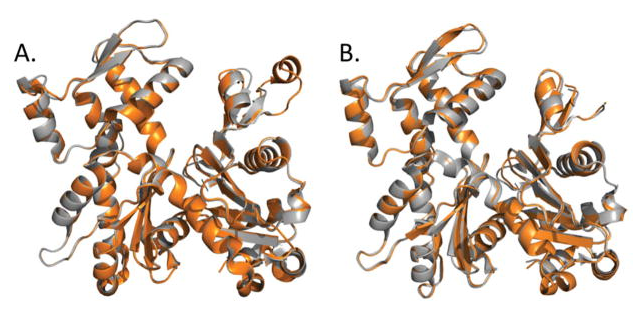
Summary: This work reviews the ATP and ADP states of a ubiquitous protein, actins, and considers the present evidence for and against unique, nucleotide-dependent conformations of this protein.
2012
25. Kudryashova E., Kalda C., Kudryashov D.S. (2012) Glutamyl phosphate is an activated intermediate in actin crosslinking by actin crosslinking domain (ACD) toxin. PLoS One 7(9):e45721 pdf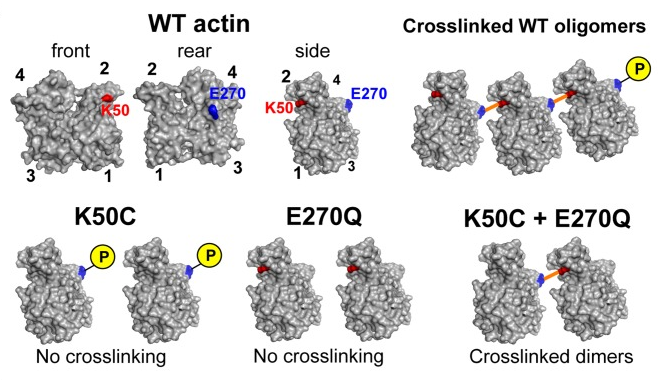
Abstract: Actin Crosslinking Domain (ACD) of MARTX toxin produced by several Gram-negative pathogenic bacteria upon delivery into the cytoplasmof eukaryotic host cells disrupts the actin cytoskeleton by catalyzing intermolecular amide bond formation between E270 and K50 residues of actin. Ultimately, accumulation of the crosslinked polymerization-deficient actin oligomers results in structural and functional failure of the actin cytoskeleton in affected cells. In the present work, we advanced in our understanding of the ACD catalytic mechanism by discovering that the enzyme transfers the gamma-phosphoryl group of ATP to the E270 actin residue, resulting in the formation of an activated acyl phosphate intermediate. This intermediate is further hydrolyzed and the energy of hydrolysis is utilized for the formation of the amide bond between actin subunits.
********************************************************************************************************************
University of California, Los Angeles
24. Oztug Durer Z.A., Kudryashov D.S., Sawaya M.R., Altenbach C., Hubbell W., Reisler E. (2012) Structural states and dynamics of the D-loop in actin. Biophysical Journal 103(5):930-9
23. Galkin V., Orlova A., Kudryashov D.S., Solodukhin A., Reisler E., Schroeder G., Egelman E. (2011) Remodeling of Actin Filaments by ADF/Cofilin Proteins. Proceedings of the National Academy of Sciences U.S.A. 108(51):20568-72.
22. Kudryashov*# D.S., Grintsevich* E.E, Rubenstein P.A., Reisler E. (2010) A Nucleotide Statesensing Region on Actin. The Journal of Biological Chemistry 285(33):25591-601.
21. Grintsevich E., Galkin V., Orlova A., Ytterberg A., Mikati M.M., Kudryashov D.S., Loo J.R., Egelman E., Reisler E. (2010) Mapping of drebrin binding site on F-actin. The Journal of Molecular Biology 498(4):542-554.
20. Durer Z., Deriviyam K., Sept D., Kudryashov D.S., Reisler E. (2010) F-Actin Structure Destabilization and DNase I Binding Loop Fluctuations Mutational Cross-Linking and Electron Microscopy Analysis of Loop States and Effects on F-Actin. The Journal of Molecular Biology 395(3):544-557.
19. Kudryashov* D.S., Durer* Z.A., Ytterberg A.J., Sawaya M.R., Pashkov I., Yeates T.0., Ogorzalek Loo R., Loo J., Satchell K.J., Reisler E. (2008) Connecting actin monomers by isopeptide bond is a toxicity mechanism of the Vibrio cholerae MARTX toxin. Proceedings of the National Academy of Sciences U.S.A. 105(47):18537-42.
This article was highlighted in: – “Research from University of California provides new data on vibrio.” Biotech Week. 2009. HighBeam Research.(January 26, 2015). http://www.highbeam.com/doc/1G1-193613844.html – (2009-02-08), Research from University of California provides new data on vibrio (Connecting actin monomers by iso-peptide bond is a toxicity mechanism of the Vibrio cholerale MARTX toxin), Health & Medicine Week, 2254, ISSN: 1532-4605, Pubz ID: 003668647. http://www.pubzpro.com/Pubz/#!search/a/3668647
18. Sawaya* M.R., Kudryashov* D.S., Pashkov* I., Reisler E., Yeates T.O. (2008) Multiple crystal structures of actin dimers and their implications for interactions in the actin filament. actin dimers. Acta Crystallographica Section D 64:454-65.
17. Kudryashov D.S., Cordero C.L., Reisler E., Satchell K.J. (2008) Characterization of the enzymatic activity of the actin cross-linking domain from the Vibrio Cholerae MART toxin. The Journal of Biological Chemistry 4;283(1):445-52.
16. Cordero C.L., Kudryashov D.S., Reisler E., Satchell K.J. (2006) The Actin cross-linking domain of the Vibrio cholerae RTX toxin directly catalyzes the covalent cross-linking of actin. The Journal of Biological Chemistry 281(43):32366-74.
15. Kudryashov# D.S., Galkin V.E., Orlova A., Phan M., Egelman E.H., Reisler E. (2006) Cofilin cross-bridges adjacent actin protomers and replaces part of the longitudinal F-actin interface. The Journal of Molecular Biology 358(3):785-97.
14. Kudryashova E., Kudryashov D.S., Kramerova I., Spencer M.J. (2005) Trim32 is a ubiquitin ligase mutated in limb girdle muscular dystrophy type 2H that binds to skeletal muscle myosin and ubiquitinates actin. The Journal of Molecular Biology 354(2):413-24.
13. Kudryashov* D.S., Sawaya* M.R., Adisetiyo H., Norcross T., Hegyi G., Reisler E., Yeates T.O. (2005) The crystal structure of a cross-linked actin dimer suggests a detailed molecular interface in F-actin. Proceedings of the National Academy of Sciences U.S.A. 102(37):13105-10.
12. Orlova A., Shvetsov A., Galkin V.E., Kudryashov D.S., Rubenstein P.A., Egelman E.H., Reisler E. (2004) Actin-destabilizing factors disrupt filaments by means of a time reversal of polymerization. Proceedings of the National Academy of Sciences U.S.A. 101(51):17664-8.
11. Muhlrad A., Kudryashov D.S., Peyser M.Y., Bobkov A.A., Almo S.C., Reisler E. (2004) Cofilin induced conformational changes in F-actin expose subdomain 2 to proteolysis. The Journal of Molecular Biology 342(5):1559-67.
10. Kudryashov D.S., Phillips M., Reisler E. (2004) Formation and destabilization of actin filaments with tetramethylrhodamine-modified actin. Biophysical Journal 87(2):1136-45.
– Editors’ choice article.
9. Kudryashov D.S., Reisler E. (2003) Solution Properties of tetramethylrhodamine modified G-actin. Biophysical Journal 85(4):2466-75.
********************************************************************************************************************
Institute of Experimental Cardiology, Russian Cardiology Research Center, Moscow
8. Kudryashov D.S., Stepanova O.V., Vilitkevich E.L., Nikonenko T.A., Nadezhdina E.S., Shanina N.A., Lukas T.J., Van Eldik L.J., Watterson D.M., Shirinsky V.P. (2004) Myosin light chain kinase (210 kDa) is a potential cytoskeleton integrator through its unique N-terminal domain. Experimental Cell Research 298(2):407-17.
7. Vilitkevich E.L., Kudryashov D.S., Stepanova O.V., Shirinsky V.P. (2004) A new actinbinding area of the myosin light chains’ high-molecular kinase. Ross. Fiziol. Zh. Im I.M. Sechenova (Rus) 90(5):577-85.
6. Kudryashov D.S., Vorotnikov A.V., Dudnakova T.V., Stepanova O.V., Lukas T.J., Sellers J.R., Watterson D.M., Shirinsky V.P. (2002) Smooth Muscle myosin filament assembly under control of a kinase-related protein (KRP) and caldesmon. The Journal of Muscle Research and Cell Motility 23(4):341-51.
5. Krymsky M.A., Kudryashov D.S., Shirinsky V.P., Lukas T.J., Watterson D.M., Vorotnikov A.V. (2001) Phosphorylation of kinase-related protein (telokin) in tonic and phasic smooth muscles. The Journal of Muscle Research and Cell Motility 22(5):425-37.
4. Vorotnikov A.V., Krymsky M.A., Chibalina M.V., Kudriashov D.S., Shirinsky V.P. (2000) Differences in contraction and regulatory protein phosphorylation of phasic and tonic smooth muscles. Tsitologia (Rus) 42:378-391.
3. Chibalina M.V., Kudriashov D.S., Shekhonin B.V., Shirinsky V.P. (2000) Functional properties and intracellular localization of high molecular weight isoform of myosin light chain kinase. Tsitologia (Rus) 42:248-252.
2. Kudryashov D.S., Chibalina M.V., Birukov K.G., Lukas T.J., Sellers J.R., Van Eldik L.J.,Watterson D.M., Shirinsky V.P. (1999) Unique sequence of a high molecular weight myosin light chain kinase is involved in interaction with actin cytoskeleton. FEBS Letters 463:67-71.
1. Bushueva T.L., Teplova M.V., Bushuev V.N., Kudriashov D.S., Vorotnikov A.V., Shirinsky V.P. (1999) Stability of the structure of KRP (kinase-related protein). Molecular Biology (Rus) 33(2):227-36.